Structure
Research on complex I structure advanced significantly in the last decade. 3D-structures of bacterial and mitochondrial enzymes (X-ray and cryo-EM) are available now in PDB:
2006 - Hydrophilic domain of respiratory complex I from Thermus thermophilus (3.3Å) 2FUG
2010 - Entire complex I from Thermus thermophilus (4.5Å) 3M9S
2010 - Membrane domain of respiratory complex I from Escherichia coli (3.9Å) 3M9C
2011 - Membrane domain of respiratory complex I from Escherichia coli (3.0Å) 3RKO
2013 - Entire complex I from Thermus thermophilus (3.3Å) 4HEA
2014 - Entire bovine heart complex I Bos taurus (4.32Å) 4UQ8
2015 - Membrane domain of bovine complex I Bos taurus (6.74Å) 5COD
2015 - Entire complex I from Yarrowia lipolytica (3.6Å) 4WZ7
2016 - Entire ovine respiratory complex I from Ovis aries (2.9Å) 5LNK
2016 - Entire respiratory complex I Bos taurus (4.3Å) class 1 5LDW and class 2 5LC5 class 3 5LDX
2018 - Entire complex I from Yarrowia lipolytica (4.32Å) 6GCS
2018 - Entire complex I (Q133C mutation in PSST subunit) from Yarrowia lipolytica (3.79Å) 6H8K
2018 - Entire complex I in the deactive state (D-form) bovine Bos taurus (4.13Å) 5O31 and mouse Mus musculus (3.9Å) 6G72
2018 - Entire complex I in the active state (A-form) mouse Mus musculus (3.3Å) 6G2J
Primary structure: Bacterial enzyme is composed of 14 subunits. Mammalian complex I is composed of at least 45 different subunits some of them are assembly cofactors. Of these, 7 are encoded by mitochondrial DNA (mtDNA) and the remaining 36 are encoded by nuclear DNA. The primary structures of the complex I subunits in many organisms is available in Uniprote (https://www.uniprot.org/)
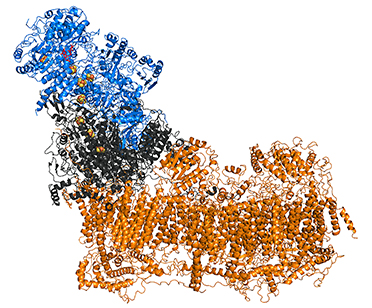
3D structure: The enzyme molecule can be divided into three functional modules (Fig on the right): peripheral N module where NADH binds (blue), central Q module that reduces ubiquinone (grey) and membrane P module (tawny) which pumps protons across the membrane (Brandt, 2016, Ann.Rev.Biochem). Complex I contains one molecule of tightly, but non-covalently bound flavin mononucleotide (FMN) and eight iron-sulfur clusters. All redox centers of the enzyme are located in the N and Q modules and there are no redox-active groups in the membrane P module.
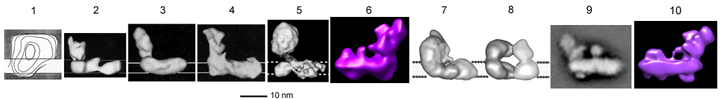
Low-resolution electron microscopy images of the enzyme from '80s and '90s are shown below.
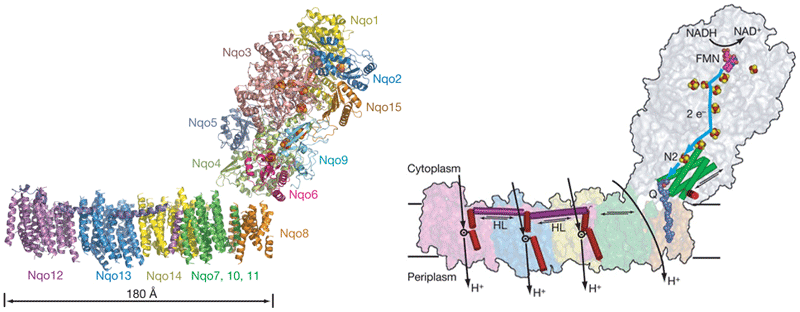
Sazanov's group first reported the structure of the entire complex I from Thermus thermophilus at 4.5 angstrom resolution. ( Efremov, R. G., Baradaran, R. and Sazanov, L. A.) *
|
|
Left: The structure consists of the atomic model for the hydrophilic domain, determined previously (PDB 3I9V), and the alpha-helical model for the membrane domain. Fe-S clusters are shown as red and yellow spheres, and FMN as magenta spheres. Each subunit is coloured differently and indicated.
Right: Proposed model of proton translocation by complex I. NADH, via FMN (magenta), donates two electrons to the chain of Fe-S clusters (red and yellow spheres), which are passed on (blue line), via terminal cluster N2, to the quinone (dark blue, moved out of the membrane by about 10 angstroms). Electron transfer is coupled to conformational changes (indicated by arrows) in the hydrophilic domain, observed for Nqo4 four-helix bundle (green cylinders) and Nqo6 helix H1 (red). These changes are transmitted to the amphipathic helix HL (magenta), which tilts three discontinuous helices (red) in antiporter-like subunits, changing the conformation of ionizable residue inside respective proton channels, resulting in translocation of three protons. The fourth proton is translocated at the interface of the two main domains.
* Reprinted by permission from Macmillan Publishers Ltd: Nature, 465, 441-445. Copyright 2010.
|
|
Mechanism of Complex I
The largest enzyme of respiratory chain complex I (or NADH:ubiquinone oxidoreductase) is a reversible redox-driven proton pump that has intrigued the bioenergetics community since the original studies from Sazanov’s group (PMID:16469879) unambiguously showed that all redox centers of the enzyme are located outside the membrane domain. Therefore, in contrast to other proton-pumping respiratory complexes, the redox centers of complex I cannot participate directly in proton translocation. Three-dimensional structures of complex I from different species have since become available, but it is still not clear how energy released during the redox reaction is transduced to drive proton pumping in the membrane domain. The operation mechanism can be divided in three interconnecting but conceptually different parts. First, the energy released during quinone binding/reduction should be converted into a conformational change in the surrounding subunits (ND1/ND3NDUFS2/NDUFS8). Second, this energy should be transferred by ~200Å, from the site of quinone reduction to distal subunit ND5. Third, this energy should drive the conformational change in the antiporter like subunits so that the protons are pumped.
The last part, i.e. the sequence of events within separate pump can be now broken down into separate molecular events resulting conformational change of the antiporter-like subunit, change in water hydration of the polar/charged residues in the broken helix element (π-bulge), and a proton is transported via a chain of water molecules by Grotthuss mechanism (see Grba and Hirst 2020, Haapanen&Sharma 2017, Di Luca et al., 2017). As suggested for the propagation of the energy via a membrane part where four successive proton pumps are located can be described as forward and reverse electrostatic waves (or wave-spring model) when first energy propagates from Q-site towards ND5 loading the pumps from one side of the membrane and then as a wave goes back protons are released on the other side (see excellent review of Kaila 2018, J. R. Soc. Interface). The main question regarding the first part however still remains - binding, reduction and release of the ubiquinone molecule is a multistep process, and it is unclear at which step energy is used to rearrange the polypeptides surrounding ubiquinone and how that is then directed towards the closest, first proton pump. Usually, some kind of interactions between complex I subunits and intermediates of quinone reduction (anionic and protonated semiquinones) were considered (see for example Brandt, 2011). Sazanov group (Gutiérrez-Fernández et al, 2020) brings a new perspective of this long-standing problem suggesting that the initial event is driven by the "tightening" of the quinone binding site and hinge motions of involved subunits to the proton translocating machinery and no redox chemistry was discussed. The reduction of the quinone (two-electron acceptor) reduction by the last FeS cluster (one-electron donor) takes place via the formation of the semiquinone first. For decades, specific semiquinone have been suggested as catalytic intermediates of Complex I, but Wright et al., 2020, reexamined the origin of semiquinone species and did not observe any semiquinone EPR signals attributed to Complex I.
Mechanistic schemes.
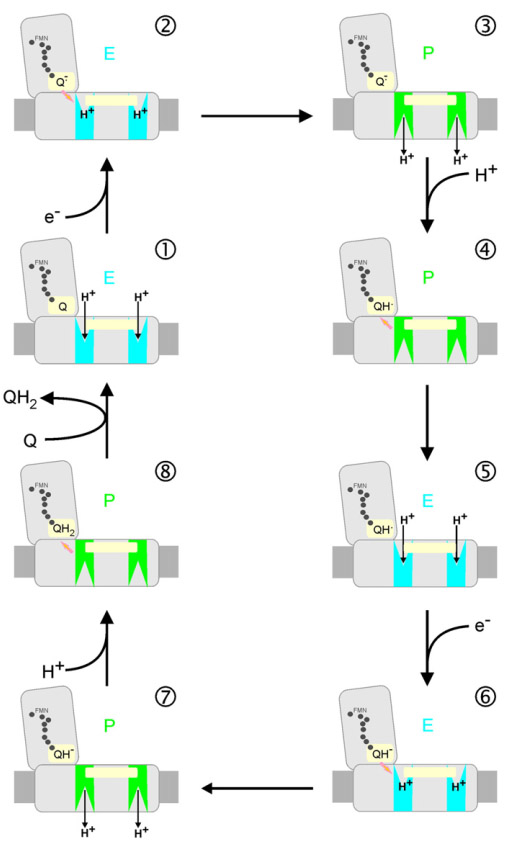
Various investigators proposed their own energy coupling mechanism of complex I (see below). These schemes were proposed before seminal Sazanov's 2006 publication (PMID:16469879) showed that there are no redox centers in the membrane domain of complex I. Currently, the most accepted hypothesis implies propagation of the conformational change from the site of the redox reactions at the Q-module to the proton pumping machinery in the membrane part. Uli Brandt proposed two-state stabilization change mechanism (PMID:21565159), in which ubiquinone redox chemistry control conformational states of the enzyme and drives proton pumping in the P module.
Figure. Pump cycle of the two-state stabilization change mechanism. The redox intermediates of ubiquinone drive conformational changes in the two pumps of the membrane P-module. The stabilization change upon formation of the anionic intermediates Q⋅− and QH− switch the pump sites from the input mode of the E-state (blue) into output mode of the P-state (green). The two pump sites may be conformationally linked by the movement of the transmission lateral helix of ∼75 residues that lies parallel to the membrane surface (light yellow bar). From PMID:21565159.
Old mechanistic schemes of complex I are below.
Ordered water molecules
In 2020 new 2.7-Å-resolution structure of complex I from aerobic yeast Yarrowia lipolytica by Grba and Hirst (PMID: 32747785) was published. This is the most accurate map of this mitochondrial enzyme to date. The structure reveals a number of ordered water molecules bound to key domains of the enzyme (Fig. 1): some supply protons to reduce ubiquinone, whereas others link critical elements of the proton-pumping machinery and probably participate directly in vectorial proton translocation. The position of some of these molecules were predicted in the bacterial enzyme experimentally and by computational approaches Efremov&Sazanov 2011, Haapanen&Sharma 2017, Di Luca et al, 2017. The position of these waters inside four proton pumps makes them direct participants in proton translocation by complex I by Grotthuss mechanism. The pump opens up by formation of water chains that are sensitive to the protonation states of buried polar and charged amino acid residues at helix elements within the pump. Watch the video here (Supplementary Video 1 from Grba and Hirst 2020, Nat.Struct.Mol.Biol).
Active/deactive transition
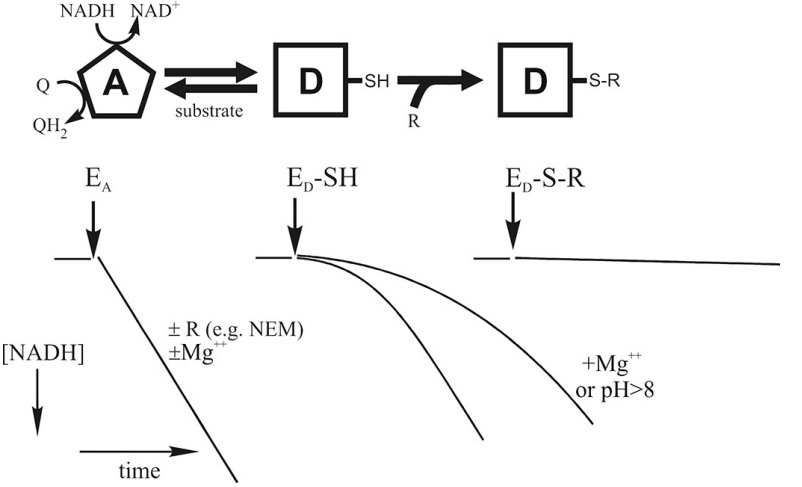
The A/D transition, which was first characterized by Andrei Vinogradov lab (PMID:2119805) for the mammalian enzyme, is also evident in other vertebrates and certain yeasts but has not been detected in prokaryotes. During steady-state respiration, the catalytically active form (A-form) is predominant and catalyzes fast reaction of oxidation of NADH by ubiquinone (10 000 min-1). If idle at physiological temperatures, i.e. during ischemia, when the lack of oxygen decreases the electron transfer in the respiratory chain, complex I converts from the A-form to the D-form. This form of the enzyme has a different conformation, it is catalytically dormant and can be reactivated during slow (1 min-1) сatalytic turnover. There is a considerable lag phase in the NADH:Q oxidoreductase reaction during reactivation of the D-form in the conventional activity essay (middle, bottom). Alkalization of the medium and divalent cations acids significantly decelerate activation.
Structural differences between the A and D-form, identified by labelling and crosslinking studies, were found to all be located close to the Q-binding site near the junction of Q and P modules (PMID:18502755, PMID:23454639, PMID:24560811, PMID:10428467). Critical thiol residue (cysteine-39) of the flexible hydrophilic loop in the membrane ND3 subunit was found to be exposed to the outside only in the D-form, but not in the A-form. If this critical cysteine residue (-SH) of complex I is modified by treatment with SH-reagents (e.g., N-ethylmaleimide) or ROS or S-nitrosothiols, then the modified enzyme (ED-S-R) is prevented from undergoing turnover-dependent reactivation.
Deactivation led to the higher exposure of several other subunits such as membrane-bound ND1 subunit and NADPH-binding subunit NDUFA9. Surprisingly, no subunits were found to be more exposed in the A-form, suggesting that the D-form corresponds to a “relaxed” conformation, while the A-form corresponds to a “tense” conformation of the enzyme. This was further confirmed by the recent cryoelectron microscopy studies of the mammalian (PMID:29395787) and yeast (PMID:3027721262) enzyme. Both catalytic forms of complex I were described for the enzyme from mouse heart (PMID:29915388) where it was revealed that significant changes occur in the transmembrane helical structures of mitochondrial-encoded ND subunits. Several catalytically important amino acids rotate upon deactivation around alpha-helical axis within the lipid bilayer. Structures of both catalytic forms are available in PDB format (PDB 6G2J and 6G72).
While it was initially characterized as purely in vitro phenomenon, A/D transition is considered one of the key events in hypoxic tissue response in the heart (PMID:23708290) and the brain (PMID:30037775, PMID:29629602). Ischemic deactivation of the enzyme would delay the return of cellular respiration back to the baseline level at the time of reoxygenation and, most likely, has a protective role
Comprehensive reviews on properties of both forms can be found here: Dröse et al 2016, PMID:26777588, Vinogradov&Grivennikova 2001 PMID:11798024, Babot et al., 2014 PMID:24569053
Evolution of Complex I (by Prof Delgi Esposti)
Complex I functions as NADH-ubiquinone (Q) oxidoreductase in many bacteria and eukaryotic mitochondria. However, this is not the original function of the ancestral forms of the enzyme complex, which has undergone biochemical transformation along with the built up of its modular structure. Read more about the evolution of the largest protonmotive enzyme complex in biology...
Inhibitors
The most well-known inhibitor specific for mitochondrial complex I is rotenone. A story about complex I inhibitor MPP+ and its link to the Parkinson's disease can be found here. However, there are a great number of compounds, both naturally-occurring and synthetic, that are potent inhibitors of complex I. Some typical inhibitors are shown in this TABLE and their structures are in this FIGURE. A comprehensive review from Delgi Esposti (PMID:9593904) covers most of the classical complex I inhibitors and new aspects and advances in this field can be found in the review from Murai and Miyoshi (PMID: 26625959).
Most of the known specific inhibitors are acting either at the beginning (NADH binding site) or the end (ubiquinone binding site) of electron transfer route in complex I. Most of hydrophobic inhibitors bind at the Q-site near FeS cluster N2 ( rotenone, piericidin A, rolliniastatin, capsaicin). Many classical detergents are also potent inhibitors of the enzyme (e.g., Triton X100) also bind there. Alternatively, arylazido-beta-alanyl NAD+, ADP-ribose, and high affinity novel inhibitor NADH-OH are competitive inhibitors that bind around NADH-binding site close to FMN.
Complex I-related diseases
As described in the Overview section, there are a number of diseases caused by defects of Complex I. Several reviews are also available from Leo Sazanov 2018 (PMID:30055843), Immo Scheffler 2015 (PMID:25224827), Jan Smeitink 2009 (PMID:19336460)
Hot topics (by Dr Volker Zickermann)
X-ray crystallography and cryoEM have unraveled the structural basis for redox-linked proton translocation by complex I. All electron transfer events take place in the peripheral arm while putative proton pump sites were found to be distributed over the whole length of the membrane arm. Ubiquinone reduction near cluster N2 triggers and drives proton pumping and structural rearrangements of the ubiquinone reduction site very likely play a key role in the coupling mechanism of complex I. However, we are just beginning to understand the catalytic cycle of complex I at the molecular level and even fundamental issues have remained unresolved. Does complex I operate by a single or dual power stroke mechanism? Which steps of ubiquinone reduction are coupled to proton translocation and how is long range energy transmission achieved? What is the role of semiquinone intermediates? What is the precise function of individual residues in the hydrophilic axis in the membrane arm? Structural information on different reaction intermediates in combination with functional studies and molecular simulation and modeling will be needed to address these questions.
Unsolved questions
1. What is the exact mechanism of Complex I energy transduction? How energy released during oxidation of terminal iron-sulfur cluster N2 located around 30Å above the membrane is converted and transferred to the membrane part where proton translocation takes place. What exactly the stages of the long-range conformational changes during catalytic turnover?
2. The fine molecular details of the A/D transition were characterized in Judy Hirst lab (PMID:29915388). Are there any similarities between the D- and the A-form and catalytic states of complex I?
3. The forward reaction of NADH oxidation by ubiquinone is reversible. Complex I can oxidize membrane ubiquinol and reduce NAD+ during reverse electron transfer. During RET, electrons are pushed “uphill” against the difference of redox potential, and this process requires a high QH2/Q ratio and the presence of a proton-motive force across the membrane. Importantly, the D to A conversion of the enzyme is only possible during the forward but not during the reverse reaction. This tells us something important about the stages of catalytic turnover.
4. Loss of FMN by mitochondrial complex I after brain ischemia results was found recently (PMID:31037949, PMID:29643256, PMID:28914132). Does it have a neuroprotective function (e.g., decrease of ROS release) or this is an initial stage of the ischemia/reperfusion injury (inactivation of the enzyme)?
5. Two tissue-specific isoforms of complex I NDUFV3 subunit was identified (PMID:27940020, PMID:27988283, PMID:27988283). Does it play any functional role and affect complex I properties and regulation in different organs?
|
