Center for Genomic Sciences. UNAM Campus de Cuernavaca
Cuernavaca, 62130 Morelos, Mexico; Email: mauro1italia@gmail.com

Complex I functions as NADH-ubiquinone (Q) oxidoreductase in many bacteria and eukaryotic mitochondria. However, this is not the original function of the ancestral forms of the enzyme complex, which has undergone biochemical transformation along the built up of its modular structure (Marreiros et al. 2013; Degli Esposti 2015; Degli Esposti 2017a). This essay summarize the evolutionary history of complex I.
The evolution of the largest protonmotive enzyme complex in biology (Sazanov 2015) has been delineated in recent studies that have examined the phylogenetic distribution of its various forms in bacterial phyla (Spero et al. 2015; Degli Esposti 2015; Marreiros et al. 2016; Degli Esposti 2017a). The proton-pumping NADH oxidoreductase of bacteria such as Thermus equally reacts with ubiquinone and menaquinone (Barabaran et al. 2013). Menaquinone (MQ, 2-methyl-3-isoprenyl-naphthalene-1,4-dione) is the dominant membrane quinone in most facultatively anaerobes and several strictly anaerobes (Degli Esposti 2017b) and has a redox potential considerable lower than that of ubiquinone.
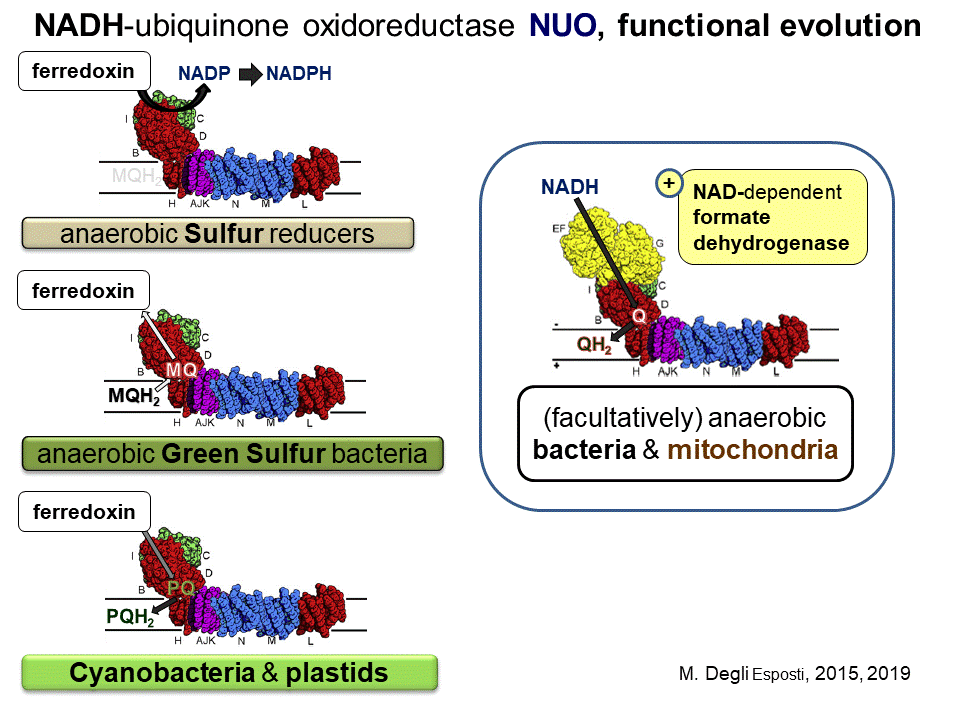
However, the most ancestral form of complex I does not react with MQ, nor Q; it transfers electrons from reduced ferredoxin to NADP+, like the Rnf complex but, most likely, without proton pumping (Degli Esposti 2015). This ancestral form of the enzyme complex is found in organisms that thrive in oceanic hydrothermal vents such as epsilonproteobacteria of the Nautiliales order (Campbell et al. 2009) and the deep branching genera of Aquificae, Desulfurobacterium and Thermovibrio (Giovannelli et al. 2012). As deduced from genomic analysis, the ancestral complex I in such strict anaerobes contains 11 subunits homologous to those of Thermus complex I (including all the transmembrane subunits), and is clearly related to membrane-bound NiFe hydrogenases (Coppi 2006; Spero et al. 2015). An hydrogenase module equivalent to that of ancestral complex I was already present in the common ancestor of all cells (Martin et al. 2016; Weiss et al. 2016), suggesting that complex I may be the oldest redox enzyme of the mitochondrial respiratory chain.
The strictly anaerobic organisms that contain the ancestral form of complex I share the pathways of H2-mediated sulfur reduction and reverse tricarboxylic acid cycle (Campbell and Cary 2004; Campbell et al. 2009), which are present also in facultatively anaerobic organisms such as Thermotoga and Magnetococcus (Schubbe et al. 2009) that possess another, partial operon for complex 1 matching that of the mbx complex originally found in the Archaean Pyrococcus (Degli Esposti, 2015). Pyrococcus mbx complex functions to support ferredoxin reducing hydrogenases, re-oxidizing reduced ferredoxin to produce NADPH for biosynthetic purposes when elementary sulfur is available as final electron acceptor (Bridger et al. 2011). The gene sequence of such a complex incorporates the entire operon of a pha/mrp antiporter system and uses the Fe–S clusters in the homolog of the NuoI subunit to directly reduce NADP+ (Degli Esposti 2015). In the mbx-like operon of proteobacteria such as Magnetococcus, the NuoI subunit is substituted by a flavoprotein related to gltD, the small subunit of NAD(P)-dependent glutamate synthase interacting with NAD(P)H. Hence, this subunit functions as the reduction site for the acceptor substrate NAD(P)+, while the mbx-like operon often terminates with rubrerythrin, a redox protein probably involved in quenching radicals (Degli Esposti 2015).
The NuoD homologs of mbx and mbx-like complexes show the vestigial presence of Cys residues at positions corresponding to cysteine ligands of the NiFe cluster of hydrogenases, a feature shared with Archaean Fpo (F420-phenazine oxidoreductase) complexes (see Degli Esposti 2015 and references therein). The same vestigial ligands are evident also in the ancestral Nuo complexes of Nautiliales, which maintain the first and third Cys ligands of the Ni–Fe cluster, corresponding to positions 86 and 403 following Thermus numeration (Drennan et al. 2001; Degli Esposti 2015). This molecular evidence is correlated with the phylogenetic sequence of the various forms of complex I based upon the NuoD subunit. Phylogenetic trees of this subunit show its homologs of Nautilales and Desulfurobacterales forming the deepest branch, followed by the homologs of the mbx-like complex and then those of the Fpo complexes, before the large clade including all the homologs from (M)Q-reacting complexes (Degli Esposti 2015).
This consistent pattern suggests that the Fpo complexes predominantly present in Archaea derived from Lateral Gene Transfer (LGT) from bacteria which had already developed the mbx-like complex, before the acquisition of quinone reactivity in the evolution of complex I (Degli Esposti 2015). Indeed, it has been documented that Archaea acquired bioenergetic enzymes via massive waves of LGT from ancestral bacteria (Nelson-Sathi et al. 2012; 2015), and either mbx or Fpo are among such enzymes. The relatively recent acquisition of respiratory enzymes in Archaean lineages, the ancestors of which were strictly anaerobes with a metabolism completely different from that of common bacteria, and mitochondria (Nelson-Sathi et al. 2012; Martin et al. 2016), is a concept that is hardly appreciated in the scientific community of Bioenergetics. However, evidence is mounting that the Archaean versions of respiratory enzymes are late comers, often highly derived forms of the ancestral enzymes originally evolved in bacterial lineages (Colman et al. 2018).
The MQ-oxidoreductase form of complex I evolved most probably in photosynthetic Chlorobi and various anaerobic bacteria, for instance Clostridiales, using ferredoxin or other low potential proteins as donor substrates (Degli Esposti 2015). Presumably, the electron flow was mainly from reduced MQ to ferredoxin, at the expense of the membrane potential generated by MQ-reducing hydrogenases. Cyanobacteria were probably the first prokaryotes to evolve the forward, protonmotive electron transport from ferredoxin to plastoquinone, a Q analog which has the same basic pathways of Q biosynthesis (Degli Esposti 2017b). This function has been thoroughly documented in recent studies with photosynthetic organisms (Schuller et al. 2019), thereby generating the name of ‘photosynthetic complex I’. The complex is formed by the same 11 core subunits as the ancestral form of the enzyme, plus additional subunits the genes of which are dispersed along the cyanobacterial (and plastidial) genome (Schuller et al. 2019).
The next stage in complex I evolution appears to have occurred in facultatively anaerobes of the deltaproteobacteria class, and led to the incorporation of the NADH-reacting subunits from soluble formate dehydrogenases (Degli Esposti 2015). Indeed, the same NADH-oxidizing subunits are retained as a separate module in hydrogenosomes and other organelles derived from mitochondria, which have lost respiratory complex I inherited from proto-mitochondria (Müller et al. 2012; Atteia et al. 2013; Degli Esposti et al. 2016).
Finally, the 14-subunit NADH-ubiquinone reductase of mitochondrial type emerged in facultatively anaerobes of the deltaproteobacterial and alphaproteobacterial class (Degli Esposti 2015), and then spread to multiple bacterial lineages possessing aerobic metabolism, presumably, again, by LGT (Spero et al. 2015). Hence, the last evolutionary stage of complex I, before being inherited by eukaryotes, is intimately connected with the development and spread of aerobic metabolism in bacteria (Degli Esposti 2017a). Once passed to the eukaryotes, the genes for the catalytic subunits of the hydrophilic arm of the complex have been progressively moved from the mtDNA derived from the bacterial symbiont to the nucleus, as recently reviewed (Degli Esposti, 2017). This survey has shown multiple situations in which the genes of some key subunits have been retained in mtDNA of protist lineages, but in a partial or deranged form that suggests loss of function (Degli Esposti, 2017). Hence, the trajectory of complex I evolution now appears to approach a parabolic pattern with progressive involution, at least in some eukaryotic lineages. The increasing number of human diseases associated with complex I defects might conform to this pattern.
References
Atteia, A., van Lis, R., Tielens, A.G. and W.F. Martin. 2013. Anaerobic energy metabolism in unicellular photosynthetic eukaryotes. Biochim. Biophys. Acta. 1827:210–223.
Baradaran, R., Berrisford, J.M., Minhas, G.S. and L.A. Sazanov. 2013. Crystal structure of the entire respiratory complex I. Nature 494: 443–448.
Bridger, S.L., Clarkson, S.M., Stirrett, K., DeBarry, M.B., Lipscomb, G.L., Schut, G.J., Westpheling, J., Scott, R.A. and M.W. Adams. 2011. Deletion strains reveal metabolic roles for key elemental sulfur-responsive proteins in Pyrococcus furiosus. J. Bacteriol. 193: 6498-6504.
Campbell, B.J. and S.C. Cary. 2004. Abundance of reverse tricarboxylic acid cycle genes in free-living microorganisms at deep-sea hydrothermal vents. Appl. Environ. Microbiol. 70: 6282-6289.
Campbell, B.J., Smith, J.L., Hanson, T.E., Klotz, M.G., Stein, L.Y. et al. 2009. Adaptations to submarine hydrothermal environments exemplified by the genome of Nautilia profundicola. PLoS Genet. 5: e1000362.
Colman, D.R., Poudel, S., Hamilton, T.L., Havig, J.R., Selensky, M.J., Shock, E.L. and E.S. Boyd. 2018. Geobiological feedbacks and the evolution of thermoacidophiles. ISME J. 12: 225-236.
Coppi, M.V. 2005. The hydrogenases of Geobacter sulfurreducens: a comparative genomic perspective. Microbiology. 151: 1239-1254.
Degli Esposti, M. 2015. Genome Analysis of Structure-Function Relationships in Respiratory Complex I, an Ancient Bioenergetic Enzyme. Genome Biol Evol. 8: 126-147. doi: 10.1093/gbe/evv239.
Degli Esposti, M., Cortez, D., Lozano, L., Rasmussen, S., Nielsen, H.B. and E. Martinez Romero. 2016. Alpha proteobacterial ancestry of the [Fe-Fe]-hydrogenases in anaerobic eukaryotes. Biol. Direct. 11: 34.
Degli Esposti, M. 2017a. The long story of mitochondrial DNA and respiratory complex I. Front Biosci (Landmark Ed). 22: 722-731.
Degli Esposti, M. 2017b. A journey across genomes uncovers the origin of ubiquinone in cyanobacteria. Genome Biol. Evol. 9: 3039-3053.
Drennan, C.L., Heo, J., Sintchak, M.D., Schreiter, E. and P.W. Ludden. 2001. Life on carbon monoxide: X-ray structure of Rhodospirillum rubrum Ni-Fe-S carbon monoxide dehydrogenase. Proc. Natl. Acad. Sci. USA. 98: 11973-11978.
Giovannelli, D., Ricci, J., Pérez-Rodríguez, I., Hügler, M., O'Brien, C., Keddis, R. et al. 2012. Complete genome sequence of Thermovibrio ammonificans HB-1(T), a thermophilic, chemolithoautotrophic bacterium isolated from a deep-sea hydrothermal vent. Stand. Genomic Sci. 7: 82-90.
Marreiros, B.C., Batista, A.P., Duarte, A.M. and M.M. Pereira. 2013. A missing link between complex I and group 4 membrane-bound [NiFe] hydrogenases. Biochim. Biophys. Acta. 1827: 198-209.
Marreiros, B.C., Calisto, F., Castro, P.J., Duarte, A.M., Sena, F.V., Silva, A.F., et al. 2016. Exploring membrane respiratory chains. Biochim. Biophys. Acta. 1857:1039-1067. doi: 10.1016/j.bbabio.2016.
Martin, W.F., Weiss, M.C., Neukirchen, S., Nelson-Sathi, S. and F.L. Sousa. 2016. Physiology, phylogeny, and LUCA. Microb. Cell. 3: 582-587. doi: 10.15698/mic2016.12.545.
Müller, M., Mentel, M., van Hellemond, J.J., Henze, K., Woehle, C., Gould, S.B., et al. 2012. Biochemistry and evolution of anaerobic energy metabolism in eukaryotes. Microbiol. Mol. Biol. Rev. 76: 444-495.
Nelson-Sathi, S., Sousa, F.L., Roettger, M., Lozada-Chávez, N., Thiergart, T., Janssen, A. et al. 2015. Origins of major archaeal clades correspond to gene acquisitions from bacteria. Nature. 517: 77-80.
Nelson-Sathi, S., Dagan, T., Landan, G., Janssen, A., Steel, M., McInerney, J.O. et al. 2012. Acquisition of 1,000 eubacterial genes physiologically transformed a methanogen at the origin of Haloarchaea. Proc. Natl. Acad. Sci. USA. 109: 20537-20542. doi: 10.1073/pnas.1209119109.
Sazanov, L.A. 2015. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nat. Rev. Mol. Cell. Biol. 16: 375-388.
Schübbe, S., Williams, T.J., Xie, G., Kiss, H.E., Brettin, T.S., Martinez, D. et al. 2009. Complete genome sequence of the chemolithoautotrophic marine magnetotactic coccus strain MC-1. Appl. Environ. Microbiol. 75: 4835-4852.
Schuller, J.M., Birrell, J.A., Tanaka, H., Konuma, T., Wulfhorst, H., Cox, N., Schuller, S.K., Thiemann, J., Lubitz, W., Sétif, P., Ikegami, T., Engel, B.D., Kurisu, G., and M.M. Nowaczyk. 2019. Structural adaptations of photosynthetic complex I enable ferredoxin-dependent electron transfer. Science. 363: 257-260.
Spero, M.A., Aylward, F.O., Currie, C.R. and T.J. Donohue. 2015. Phylogenomic analysis and predicted physiological role of the proton-translocating NADH:quinone oxidoreductase (complex I) across bacteria. MBio. 6:ve00389-15.
Weiss, M.C., Sousa, F.L., Mrnjavac, N., Neukirchen, S., Roettger, M., Nelson-Sathi, S. et al. The physiology and habitat of the last universal common ancestor. Nat. Microbiol. 1:16116. doi:10.1038/nmicrobiol.2016.116.